Active learning framework leveraging transcriptomics identifies modulators of disease phenotypes | Science
Phenotypic drug screening, a process integral to discovering new therapeutics, faces significant challenges due to the immense diversity of chemical compounds and the complexity of scaling experimental workflows. The traditional approach, which involves testing a wide array of compounds on biological systems to observe phenotypic changes, is often hampered by logistical and resource constraints. As the number of potential compounds grows, the ability to efficiently identify promising candidates becomes increasingly difficult. To address these issues, researchers have begun to adopt computational methods that prioritize compounds based on predicted activity and biological relevance. These methods aim to streamline the screening process, making it more efficient and less resource-intensive.
However, the reliance on computational predictions introduces its own set of challenges. Many of these methods depend on existing biological data, which can be limited or biased. For instance, if the training data used to develop predictive models lacks diversity or is not representative of the chemical space, the predictions may not accurately reflect the true potential of untested compounds. This can lead to missed opportunities for discovering novel therapeutics. Furthermore, the integration of computational approaches with experimental workflows poses technical hurdles, as researchers must ensure that the predictions align with real-world biological responses. Despite these challenges, the ongoing development of more sophisticated algorithms and machine learning techniques holds promise for enhancing the efficacy of phenotypic drug screening.
Recent advancements in computational methods have shown potential in overcoming these barriers. For example, deep learning techniques have been employed to analyze large datasets, uncovering hidden patterns that traditional methods might overlook. Additionally, the incorporation of multi-omics data—combining genomics, proteomics, and metabolomics—can provide a more holistic view of biological responses, enabling more accurate predictions of compound efficacy. As these technologies continue to evolve, they may significantly enhance the ability to navigate the vast chemical space, ultimately leading to more effective drug discovery processes. In summary, while phenotypic drug screening is currently constrained by various factors, the integration of computational methods offers a pathway to improve efficiency and accuracy in identifying promising drug candidates.
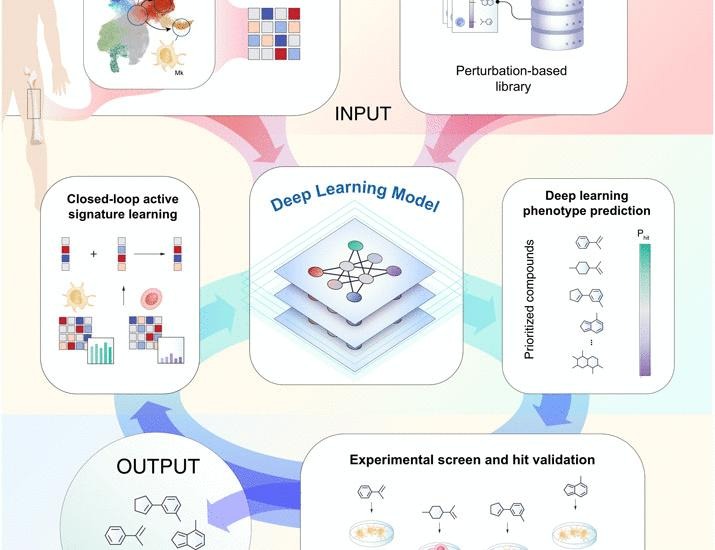
Phenotypic drug screening remains constrained by the vastness of chemical space and the technical challenges of scaling experimental workflows. To overcome these barriers, computational methods have been developed to prioritize compounds, but they rely …