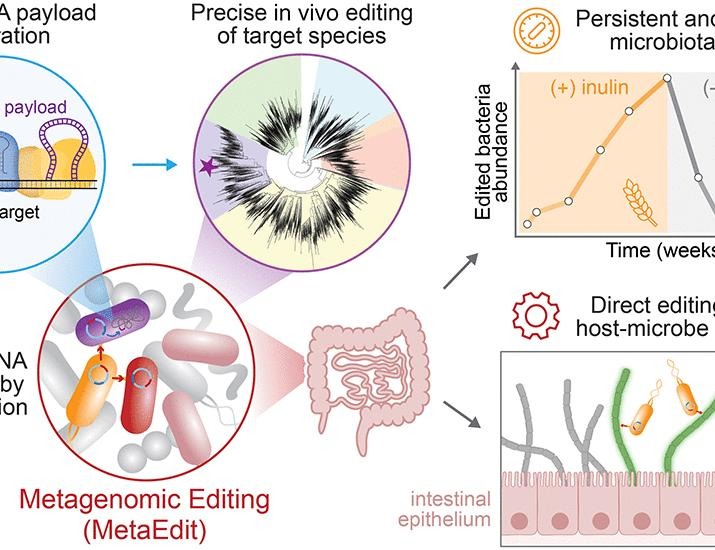
Metagenomic editing of commensal bacteria in vivo using CRISPR-associated transposases | Science
**Revolutionizing Microbiome Research: The Introduction of Metagenomic Editing (MetaEdit)**
In recent years, metagenomic sequencing has unveiled the vast and complex biodiversity of microbial communities residing in the mammalian gut. These microbial populations play crucial roles in various physiological processes, including digestion, immune function, and even mental health. However, despite the wealth of information regarding these microbial species, the ability to genetically manipulate specific bacteria within the microbiome has remained a significant challenge. Traditional genetic editing techniques, such as CRISPR, have been limited in their application to the gut microbiome due to the intricate interactions between microbial species and the difficulty in targeting specific organisms without affecting others.
To address this gap, researchers have introduced a groundbreaking platform known as Metagenomic Editing (MetaEdit). This innovative approach allows for the precise alteration of genetic material within specific microbial species in the gut, paving the way for targeted therapies and enhanced understanding of microbial functions. MetaEdit leverages advanced sequencing technologies and bioinformatics tools to identify and modify genes of interest within the microbiome, facilitating the study of their roles in health and disease. For example, by selectively editing genes linked to metabolic pathways, scientists can explore how changes in microbial composition affect the host’s metabolism and overall health.
The implications of MetaEdit are profound, as it opens up new avenues for research and therapeutic interventions. For instance, by targeting specific gut bacteria associated with conditions like obesity or inflammatory bowel disease, researchers could develop tailored treatments that restore a healthy microbial balance. Furthermore, MetaEdit could enhance our understanding of the gut-brain axis, where gut microbiota influence neurological functions. As this technology continues to evolve, it promises to transform our approach to microbiome research, providing deeper insights into the intricate relationships between humans and their microbial inhabitants. The potential applications of MetaEdit could lead to significant advancements in personalized medicine and the treatment of various diseases linked to gut microbiota.
Although metagenomic sequencing has revealed a rich microbial biodiversity in the mammalian gut, methods to genetically alter specific species in the microbiome are highly limited. Here, we introduce Metagenomic Editing (MetaEdit) as a platform …